1010Genome offers next generation sequencing data analysis through Illumina sequencing platform. The cost effective, optimized solutions for whole exome sequencing are critical for complex disease studies to discover SNP biomarkers. Similarly, Illumina sequencing is a very time and cost effective platform for gene expression in eukaryotes. Illumina sequencing has been acknowledged as one among the leading and accurate methods for generating readings that help in the comparison of DNA sequences. Millions of reads that are quickly and accurately generated by this method are aligned to a reference genome or transcriptome for comparison. Two standout advantages of this method include the high speed with which the reads are generated and the low cost of the method.

Illumina Sequencing Instruments from desktop to production.
Overview of Illumina Sequencing
- Illumina sequencing library preparation starts with breaking down gDNA into shorter fragments.
- These short fragments are then ligated to adapters at both ends.
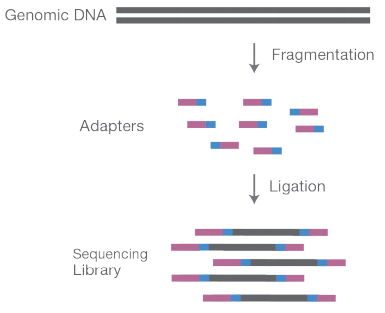
3. Library is then loaded on to a flow cell for cluster amplification process.
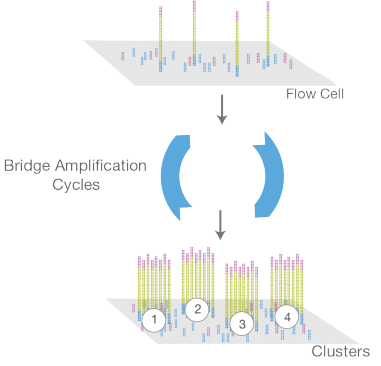
4. Flow cell is lined with oligos complimentary to adapters resulting into a hybridization of library.
5. This is followed by a bridge amplification cycles that generates clusters.
6. These clusters are colonies that originate from a single library fragment. It is important to generate such clusters as fluorescence from a single library molecule would go undetected.
7. Flurocently labeled nucleotide bases and sequencing reagents are then added for first base incorporation reaction.
8. Next, a specific wavelength and intensity is used to detect and capture fluorescence from each of cluster to detect bases.
 9. This cycle of adding bases and image capture is followed for the desired length of reads. Example 100 bp or 150 bps.
9. This cycle of adding bases and image capture is followed for the desired length of reads. Example 100 bp or 150 bps.
10. Captured images are processed by sophisticated software and then output as read bases in base call files – bcl or converted into fasta files that are text files.

11. These reads are now aligned to a reference genome or transcriptome for mapping and further analysis.


