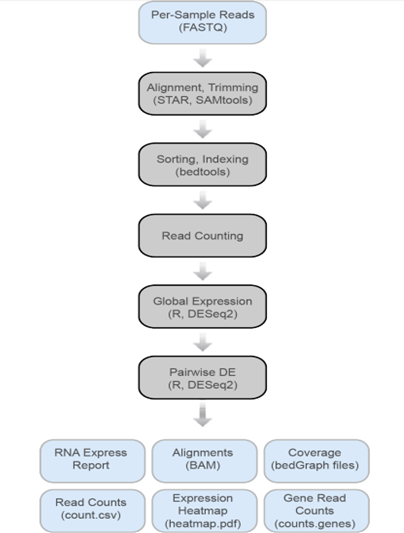
Differential Gene Expression Tools
Welcome to 1010Genome's comprehensive guide on analyzing differential gene expression using RNA-Seq data. In this guide, we will introduce you to the exciting field of bioinformatics and provide you with an overview of the tools and resources available to help you unlock the mysteries of gene expression.
Understanding RNA-Seq and Differential Gene Expression
RNA-Seq (RNA sequencing) is a cutting-edge technology that allows us to investigate the transcriptome of an organism. It provides a detailed snapshot of which genes are actively being transcribed and their expression levels at a particular moment in time. Differential gene expression analysis is the process of comparing the transcriptomes of different biological conditions or samples to identify genes that are differentially expressed. Studying differential gene expression is crucial for understanding how genes respond to various conditions, such as diseases, treatments, or environmental changes.

Best Practices for RNA-Seq Differential Gene Expression Analysis
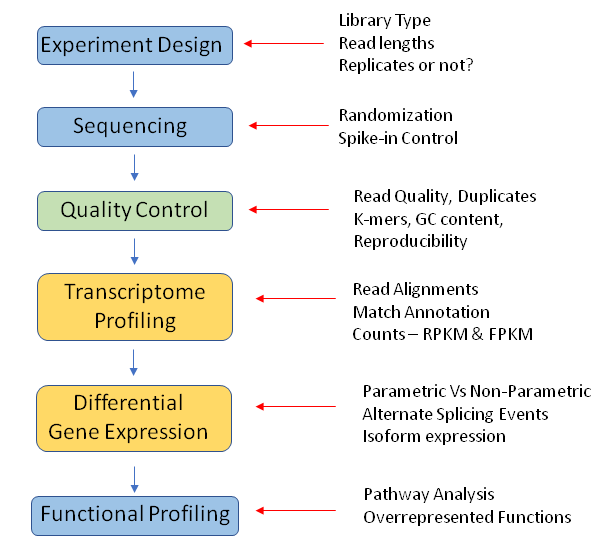
Successful RNA-Seq differential gene expression analysis requires careful planning and execution. Here are some best practices to keep in mind:
- Experimental Design: Plan your experiment carefully, ensuring proper sample selection, biological replicates, and control groups.
- Quality Control: Regularly check the quality of your data at each step, from raw sequencing to the final analysis.
- Normalization: Apply appropriate normalization methods to account for technical variations in your data.
- Statistical Tests: Use appropriate statistical tests to identify differentially expressed genes.
- Data Visualization: Visualize your results to gain insights and validate your findings.
- Functional Annotation: Use database resources to understand the biological context of your results.
Tools for Differential Gene Expression Analysis
At 1010Genome, we offer a suite of powerful bioinformatics tools designed to facilitate the analysis of RNA-Seq data for differential gene expression.

1. FastQC: Quality Control
Before diving into differential gene expression analysis, it’s essential to check the quality of your raw sequencing data. FastQC is a tool that helps you assess the quality of your data by generating various quality control metrics and reports. This ensures that your analysis starts with reliable data.
2. STAR and HISAT2: Aligning Sequences
STAR and HISAT2 are popular alignment tools that map the sequencing reads to a reference genome. These tools ensure that the RNA-Seq reads are accurately aligned, setting the stage for downstream analysis.
3. FeatureCounts and HTSeq: Counting Reads
Once your reads are aligned, you need to count how many reads are associated with each gene. FeatureCounts and HTSeq are tools that perform this critical step in the analysis, generating count tables for downstream analysis.
4. DESeq2 and edgeR: Differential Expression Analysis
DESeq2 and edgeR are two widely used tools for performing differential gene expression analysis. These tools compare the read counts between different conditions or groups and identify genes that are significantly differentially expressed. They help you understand which genes play a crucial role in the observed biological differences.
5. Visualization Tools
Exploring the results of your differential gene expression analysis is crucial. Tools like R (with ggplot2 and other visualization packages), Python (with Matplotlib and Seaborn), and interactive tools like Shiny can be employed to generate informative plots and visualizations to aid in interpretation.
6. Database Resources
1010Genome provides access to comprehensive databases, including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG), which can help you functionally annotate and interpret your differentially expressed genes.
Get Started with Differential Gene Expression Analysis
Ready to start your journey into the world of RNA-Seq and differential gene expression analysis? 1010Genome is here to support you with a range of tools, resources, and expert guidance. Whether you're a novice or an experienced bioinformatician, our platform is designed to help you make the most of your RNA-Seq data.